|
Amyloid
Precursor Protein (APP)
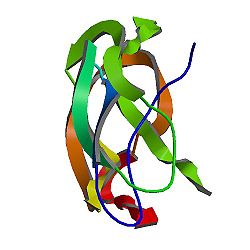 |
Amyloid
beta (A4) precursor protein (peptidase nexin-II, Alzheimer
disease)
PDB
rendering based on 1aap
|
Amyloid
precursor protein (APP) is an integral membrane protein expressed
in many tissues and concentrated in the synapses of neurons.
Its primary function is not known, though it has been implicated
as a regulator of synapse formation[2] and neural plasticity.[3]
APP is best known and most commonly studied as the precursor
molecule whose proteolysis generates amyloid beta (Aß), a 39-
to 42-amino acid peptide whose amyloid fibrillar form is the
primary component of amyloid plaques found in the brains of
Alzheimer's disease patients.
Genetics
In
humans, the gene for APP is located on chromosome 21 and contains
at least 18 exons in 240 kilobases.[4][5] Several alternative
splicing isoforms of APP have been observed in humans, ranging
in length from 365 to 770 amino acids, with certain isoforms
preferentially expressed in neurons; changes in the neuronal
ratio of these isoforms have been associated with Alzheimer's
disease.[6] Homologous proteins have been identified in other
organisms such as Drosophila (fruit flies), C. elegans (roundworms),
and all mammals.[7] The amyloid beta region of the protein,
located in the membrane-spanning domain, is not well conserved
across species and has no obvious connection with APP's native-state
biological functions.[7]
Mutations
in critical regions of Amyloid Precursor Protein, including
the region that generates amyloid beta, are known to cause familial
susceptibility to Alzheimer's disease.[8][9][10] For example,
several mutations outside the Aß region associated with familial
Alzheimer's have been found to dramatically increase production
of Aß.[11]
Structure
A
number of distinct, largely independently-folding structural
domains have been identified in the APP sequence. The extracellular
region, much larger than the intracellular region, is divided
into the E1 and E2 domains; E1 contains several subdomains including
a growth factor-like domain (GFLD), a metal-binding motif, and
a serine protease inhibitor domain that is absent from the isoform
differentially expressed in the brain.[12] The E2 domain contains
a coiled coil dimerization motif and may bind proteoglycans
in the extracellular matrix.[1]
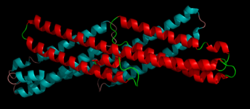 |
The
extracellular E2 domain, a dimeric coiled coil and one of
the most highly-conserved regions of the protein from Drosophila
to humans. This domain, which resembles the structure of
spectrin, is thought to bind heparan sulfate proteoglycans.[1] |
The
complete crystal structure of APP has not yet been solved; however,
individual domains have been successfully crystallized, including
the copper-binding as well as a zinc-binding domain in multiple
configurations and ion-binding states[13] and the E2 dimerization
domain.[1]
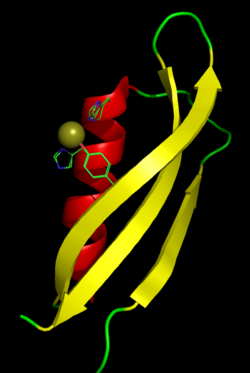 |
The
metal-binding domain of APP with a bound copper ion. The
side chains of the two histidine and one tyrosine residues
that play a role in metal coordination are shown in the
Cu(I) bound, Cu(II) bound, and unbound conformations, which
differ by only small changes in orientation. |
Post-translational
processing
APP
undergoes extensive post-translational modification including
glycosylation, phosphorylation, and tyrosine sulfation, as well
as many types of proteolytic processing to generate peptide
fragments.[14] It is commonly cleaved by proteases in the secretase
family; alpha secretase and beta secretase both remove nearly
the entire extracellular domain to release membrane-anchored
carboxy-terminal fragments that may be associated with apoptosis.[7]
Cleavage by gamma secretase within the membrane-spanning domain
generates the amyloid-beta fragment; gamma secretase is a large
multi-subunit complex whose components have not yet been fully
characterized, but include presenilin, whose gene has been identified
as a major genetic risk factor for Alzheimer's.[15] The amyloidogenic
processing of APP has been linked to its presence in lipid rafts.
When APP molecules occupy a lipid raft region of membrane, they
are more accessible to and differentially cleaved by beta secretase,
whereas APP molecules outside a raft are differentially cleaved
by the non-amyloidogenic alpha secretase.[16] Gamma secretase
activity has also been associated with lipid rafts.[17] The
role of cholesterol in lipid raft maintenance has been cited
as a likely explanation for observations that high cholesterol
and apolipoprotein E genotype are major risk factors for Alzheimer's
disease.[18]
Biological
function
Although
the native biological role of APP is of obvious interest to
Alzheimer's research, thorough understanding has remained elusive.
The most-substantiated role for APP is in synaptic formation
and repair;[2] its expression is upregulated during neuronal
differentiation and after neural injury. Roles in cell signaling,
long-term potentiation, and cell adhesion have been proposed
and supported by as-yet limited research.[7] In particular,
similarities in post-translational processing have invited comparisons
to the signaling role of the surface receptor protein Notch.[19]
APP knockout mice are viable and have relatively minor phenotypic
effects including impaired long-term potentiation and memory
loss without general neuron loss.[20] On the other hand, transgenic
mice with upregulated APP expression have also been reported
to show impaired long-term potentiation.[21] The logical inference
is that because Aß accumulates excessively in Alzheimer's disease
its precursor, APP, would be elevated as well. However, neuronal
cell bodies contain less APP as a function of their proximity
to amyloid plaques.[22] The data indicate that this deficit
in APP results from a decline in production rather than an increase
in catalysis. Loss of a neuron's APP may effect physiological
deficits that contribute to dementia.
Interactions
Amyloid
precursor protein has been shown to interact with
APBA3,[23][24]
CLSTN1,[25][26]
APPBP1,[27]
Gelsolin,[28]
BCAP31,[29]
Caveolin
1,[30]
FBLN1,[31]
Collagen, type
XXV, alpha 1,[32]
APBB1,[33][34][35][36][37]
APBA2,[23][26][38]
APBA1,[23][33]
APPBP2,[39]
HSD17B10,[40]
BLMH[41]
and SHC1.[42]
One groups
of scientists reports that APP interacts with reelin, a protein implicated in
a number of brain disorders, including Alzheimer's disease.[43]
References
1
a b c Wang Y, Ha Y. (2006). The X-ray structure of an antiparallel
dimer of the human amyloid precursor protein E2 domain. Mol
Cell 15(3):343-53. PMID 15304215
2
a b Priller C, Bauer T, Mitteregger G, Krebs B, Kretzschmar
HA, Herms J. (2006). Synapse formation and function is modulated
by the amyloid precursor protein. J Neurosci 26(27):7212-21.
PMID 16822978
3
Turner PR, O'Connor K, Tate WP, Abraham WC. (2003). Roles of
amyloid precursor protein and its fragments in regulating neural
activity, plasticity, and memory. Prog Neurobiol 70(1):1-32.
PMID 12927332
4
Yoshikai S, Sasaki H, Doh-ura K, Furuya H, Sakaki Y (1990).
Genomic organization of the human amyloid beta-protein precursor
gene Gene 87:257-263. PMID 2110105
5
Lamb BT, Sisodia SS, Lawler AM, Slunt HH, Kitt CA, Kearns WG,
Pearson PL, Price DL, Gearhart JD. (1993). Introduction and
expression of the 400 kilobase amyloid precursor protein gene
in transgenic mice Nat Genet 5:22-30. PMID 8220418
6
Matsui T, Ingelsson M, Fukumoto H, Ramasamy K, Kowa H, Frosch
MP, Irizarry MC, Hyman BT. (2007). Expression of APP pathway
mRNAs and proteins in Alzheimer's disease. Brain Res Epub. PMID
17586478
7
a b c d Zheng H, Koo EH. (2006). The amyloid precursor protein:
beyond amyloid. Mol Neurodegener 3;1:5. PMID 16930452
8
Goate A, Chartier-Harlin MC, Mullan M, Brown J, Crawford F,
Fidani L, Giuffra L, Haynes A, Irving N, James L, et al. (1991).
Segregation of a missense mutation in the amyloid precursor
protein gene with familial Alzheimer's disease. Nature 349(6311):704-6.
PMID 1671712
9
Murrell J, Farlow M, Ghetti B, Benson MD. (1991). A mutation
in the amyloid precursor protein associated with hereditary
Alzheimer's disease. Science 254(5028):97-9. PMID 1925564
10
Chartier-Harlin MC, Crawford F, Houlden H, Warren A, Hughes
D, Fidani L, Goate A, Rossor M, Roques P, Hardy J, et al. (1991).
Early-onset Alzheimer's disease caused by mutations at codon
717 of the beta-amyloid precursor protein gene. Nature 353(6347):844-6.
PMID 1944558
11
Citron M, Oltersdorf T, Haass C, McConlogue L, Hung AY, Seubert
P, Vigo-Pelfrey C, Lieberburg I, Selkoe DJ. (1992). Mutation
of the beta-amyloid precursor protein in familial Alzheimer's
disease increases beta-protein production. Nature 360(6405):672-4.
PMID 1465129
12
Sisodia SS, Koo EH, Hoffman PN, Perry G, Price DL. (1993). Identification
and transport of full-length amyloid precursor proteins in rat
peripheral nervous system. J Neurosci 13:3136-3142. PMID 8331390
13
Kong GK, Galatis D, Barnham KJ, Polekhina G, Adams JJ, Masters
CL, Cappai R, Parker MW, McKinstry WJ. (2005). Crystallization
and preliminary crystallographic studies of the copper-binding
domain of the amyloid precursor protein of Alzheimer's disease.
Acta Crystallograph 61(Pt 1):93-5. PMID 16508101. See also 2007
PDB IDs 2FJZ, 2FK2, 2FKL.
14
De Strooper B, Annaert W. (2000). Proteolytic processing and
cell biological functions of the amyloid precursor protein.
J Cell Sci 113 ( Pt 11):1857-70. PMID 10806097
15
Chen F, Hasegawa H, Schmitt-Ulms G, Kawarai T, Bohm C, Katayama
T, Gu Y, Sanjo N, Glista M, Rogaeva E, Wakutani Y, Pardossi-Piquard
R, Ruan X, Tandon A, Checler F, Marambaud P, Hansen K, Westaway
D, St George-Hyslop P, Fraser P. (2006). TMP21 is a presenilin
complex component that modulates gamma-secretase but not epsilon-secretase
activity. Nature 440:1208-1212. PMID 16641999
16
Ehehalt R, Keller P, Haass C, Thiele C, Simons K. (2003). Amyloidogenic
processing of the Alzheimer beta-amyloid precursor protein depends
on lipid rafts. J Cell Biol 160(1):113-23. PMID 12515826
17
Vetrivel KS, Cheng H, Lin W, Sakurai T, Li T, Nukina N, Wong
PC, Xu H, Thinakaran G. (2004). Association of gamma-secretase
with lipid rafts in post-Golgi and endosome membranes. J Biol
Chem 279(43):44945-54. PMID 15322084
18
Riddell DR, Christie G, Hussain I, Dingwall C. (2001). Compartmentalization
of beta-secretase (Asp2) into low-buoyant density, noncaveolar
lipid rafts. Curr Biol 11(16):1288-93. PMID 11525745
19
Selkoe D, Kopan R. (2003). Notch and Presenilin: regulated intramembrane
proteolysis links development and degeneration. Annu Rev Neurosci
26:565-597. PMID 12730322
20
Phinney AL, Calhoun ME, Wolfer DP, Lipp HP, Zheng H, Jucker
M. (1999). No hippocampal neuron or synaptic bouton loss in
learning-impaired aged beta-amyloid precursor protein-null mice.
Neuroscience 90(4):1207-16. PMID 10338291
21
Matsuyama S, Teraoka R, Mori H, Tomiyama T. (2007). Inverse
correlation between amyloid precursor protein and synaptic plasticity
in transgenic mice. Neuroreport 18(10):1083-7. PMID 17558301
22
Barger SW, DeWall KM, Liu L, Mrak RE, Griffin WS (August 2008).
"Relationships between expression of apolipoprotein E and beta-amyloid
precursor protein are altered in proximity to Alzheimer beta-amyloid
plaques: potential explanations from cell culture studies".
J. Neuropathol. Exp. Neurol. 67 (8): 773–83. doi:10.1097/NEN.0b013e318180ec47.
PMID 18648325.
23
a b c Biederer, Thomas; Cao Xinwei, Südhof Thomas C, Liu Xinran
(Sep. 2002). "Regulation of APP-dependent transcription complexes
by Mint/X11s: differential functions of Mint isoforms". J. Neurosci.
(United States) 22 (17): 7340-51. PMID 12196555.
24
Tanahashi, H; Tabira T (Feb. 1999). "X11L2, a new member of
the X11 protein family, interacts with Alzheimer's beta-amyloid
precursor protein". Biochem. Biophys. Res. Commun. (UNITED STATES)
255 (3): 663-7. doi:10.1006/bbrc.1999.0265. ISSN 0006-291X.
PMID 10049767.
25
Araki, Yoichi; Miyagi Naomi, Kato Naoko, Yoshida Tomohiro, Wada
Sachiyo, Nishimura Masaki, Komano Hiroto, Yamamoto Tohru, De
Strooper Bart, Yamamoto Kazuo, Suzuki Toshiharu (Jun. 2004).
"Coordinated metabolism of Alcadein and amyloid beta-protein
precursor regulates FE65-dependent gene transactivation". J.
Biol. Chem. (United States) 279 (23): 24343-54. doi:10.1074/jbc.M401925200.
ISSN 0021-9258. PMID 15037614.
26
b Araki, Yoichi; Tomita Susumu, Yamaguchi Haruyasu, Miyagi Naomi,
Sumioka Akio, Kirino Yutaka, Suzuki Toshiharu (Dec. 2003). "Novel
cadherin-related membrane proteins, Alcadeins, enhance the X11-like
protein-mediated stabilization of amyloid beta-protein precursor
metabolism". J. Biol. Chem. (United States) 278 (49): 49448-58.
doi:10.1074/jbc.M306024200. ISSN 0021-9258. PMID 12972431.
27
Chow, N; Korenberg J R, Chen X N, Neve R L (May. 1996). "APP-BP1,
a novel protein that binds to the carboxyl-terminal region of
the amyloid precursor protein". J. Biol. Chem. (UNITED STATES)
271 (19): 11339-46. ISSN 0021-9258. PMID 8626687.
28
Chauhan, V P; Ray I, Chauhan A, Wisniewski H M (May. 1999).
"Binding of gelsolin, a secretory protein, to amyloid beta-protein".
Biochem. Biophys. Res. Commun. (UNITED STATES) 258 (2): 241-6.
doi:10.1006/bbrc.1999.0623. ISSN 0006-291X. PMID 10329371.
29
Wang, Bing; Nguyen Mai, Breckenridge David G, Stojanovic Marina,
Clemons Paul A, Kuppig Stephan, Shore Gordon C (Apr. 2003).
"Uncleaved BAP31 in association with A4 protein at the endoplasmic
reticulum is an inhibitor of Fas-initiated release of cytochrome
c from mitochondria". J. Biol. Chem. (United States) 278 (16):
14461-8. doi:10.1074/jbc.M209684200. ISSN 0021-9258. PMID 12529377.
30
Ikezu, T; Trapp B D, Song K S, Schlegel A, Lisanti M P, Okamoto
T (Apr. 1998). "Caveolae, plasma membrane microdomains for alpha-secretase-mediated
processing of the amyloid precursor protein". J. Biol. Chem.
(UNITED STATES) 273 (17): 10485-95. ISSN 0021-9258. PMID 9553108.
31
Ohsawa, I; Takamura C, Kohsaka S (Mar. 2001). "Fibulin-1 binds
the amino-terminal head of beta-amyloid precursor protein and
modulates its physiological function". J. Neurochem. (United
States) 76 (5): 1411-20. ISSN 0022-3042. PMID 11238726.
32
Hashimoto, Tadafumi; Wakabayashi Tomoko, Watanabe Atsushi, Kowa
Hisatomo, Hosoda Ritsuko, Nakamura Atsushi, Kanazawa Ichiro,
Arai Takao, Takio Koji, Mann David M A, Iwatsubo Takeshi (Apr.
2002). "CLAC: a novel Alzheimer amyloid plaque component derived
from a transmembrane precursor, CLAC-P/collagen type XXV". EMBO
J. (England) 21 (7): 1524-34. doi:10.1093/emboj/21.7.1524. ISSN
0261-4189. PMID 11927537.
33
a b Borg, J P; Ooi J, Levy E, Margolis B (Nov. 1996). "The phosphotyrosine
interaction domains of X11 and FE65 bind to distinct sites on
the YENPTY motif of amyloid precursor protein". Mol. Cell. Biol.
(UNITED STATES) 16 (11): 6229-41. ISSN 0270-7306. PMID 8887653.
34
Zambrano, N; Buxbaum J D, Minopoli G, Fiore F, De Candia P,
De Renzis S, Faraonio R, Sabo S, Cheetham J, Sudol M, Russo
T (Mar. 1997). "Interaction of the phosphotyrosine interaction/phosphotyrosine
binding-related domains of Fe65 with wild-type and mutant Alzheimer's
beta-amyloid precursor proteins". J. Biol. Chem. (UNITED STATES)
272 (10): 6399-405. ISSN 0021-9258. PMID 9045663.
35
Guénette, S Y; Chen J, Jondro P D, Tanzi R E (Oct. 1996). "Association
of a novel human FE65-like protein with the cytoplasmic domain
of the beta-amyloid precursor protein". Proc. Natl. Acad. Sci.
U.S.A. (UNITED STATES) 93 (20): 10832-7. ISSN 0027-8424. PMID
8855266.
36
Tanahashi, H; Tabira T (Feb. 1999). "Molecular cloning of human
Fe65L2 and its interaction with the Alzheimer's beta-amyloid
precursor protein". Neurosci. Lett. (IRELAND) 261 (3): 143-6.
ISSN 0304-3940. PMID 10081969.
37
Trommsdorff, M; Borg J P, Margolis B, Herz J (Dec. 1998). "Interaction
of cytosolic adaptor proteins with neuronal apolipoprotein E
receptors and the amyloid precursor protein". J. Biol. Chem.
(UNITED STATES) 273 (50): 33556-60. ISSN 0021-9258. PMID 9837937.
38
Tomita, S; Ozaki T, Taru H, Oguchi S, Takeda S, Yagi Y, Sakiyama
S, Kirino Y, Suzuki T (Jan. 1999). "Interaction of a neuron-specific
protein containing PDZ domains with Alzheimer's amyloid precursor
protein". J. Biol. Chem. (UNITED STATES) 274 (4): 2243-54. ISSN
0021-9258. PMID 9890987.
39
Zheng, P; Eastman J, Vande Pol S, Pimplikar S W (Dec. 1998).
"PAT1, a microtubule-interacting protein, recognizes the basolateral
sorting signal of amyloid precursor protein". Proc. Natl. Acad.
Sci. U.S.A. (UNITED STATES) 95 (25): 14745-50. ISSN 0027-8424.
PMID 9843960.
40
Yan, S D; Fu J, Soto C, Chen X, Zhu H, Al-Mohanna F, Collison
K, Zhu A, Stern E, Saido T, Tohyama M, Ogawa S, Roher A, Stern
D (Oct. 1997). "An intracellular protein that binds amyloid-beta
peptide and mediates neurotoxicity in Alzheimer's disease".
Nature (ENGLAND) 389 (6652): 689-95. doi:10.1038/39522. ISSN
0028-0836. PMID 9338775.
41
Lefterov, I M; Koldamova R P, Lazo J S (Sep. 2000). "Human bleomycin
hydrolase regulates the secretion of amyloid precursor protein".
FASEB J. (UNITED STATES) 14 (12): 1837-47. ISSN 0892-6638. PMID
10973933.
42
Tarr, Philip E; Roncarati Roberta, Pelicci Giuliana, Pelicci
Pier Giuseppe, D'Adamio Luciano (May. 2002). "Tyrosine phosphorylation
of the beta-amyloid precursor protein cytoplasmic tail promotes
interaction with Shc". J. Biol. Chem. (United States) 277 (19):
16798-804. doi:10.1074/jbc.M110286200. ISSN 0021-9258. PMID
11877420.
43
Study paper, a review at Alzheimer's Research Forum, and a news
report: Hoe HS, Lee KJ, Carney RS, Lee J, Markova A, Lee JY,
Howell BW, Hyman BT, Pak DT, Bu G, Rebeck GW (June 2009). "Interaction
of reelin with amyloid precursor protein promotes neurite outgrowth".
J. Neurosci. 29 (23): 7459–73. doi:10.1523/JNEUROSCI.4872-08.2009.
PMID 19515914. http://www.jneurosci.org/cgi/pmidlookup?view=long&pmid=19515914.
"Another Take on APP and Neurite Outgrowth—The Role of Reelin
(Tom Fagan, 19 June 2009)". http://www.alzforum.org/new/detail.asp?id=2170.
Retrieved 2009-06-24. "Protein linked to Alzheimer's disease
doesn't act alone (www.sciencecodex.com)". http://www.sciencecodex.com/protein_linked_to_alzheimers_disease_doesnt_act_alone.
Retrieved 2009-06-16.
|

